List of research infrastructures
Infrastructure resources at Lund University.
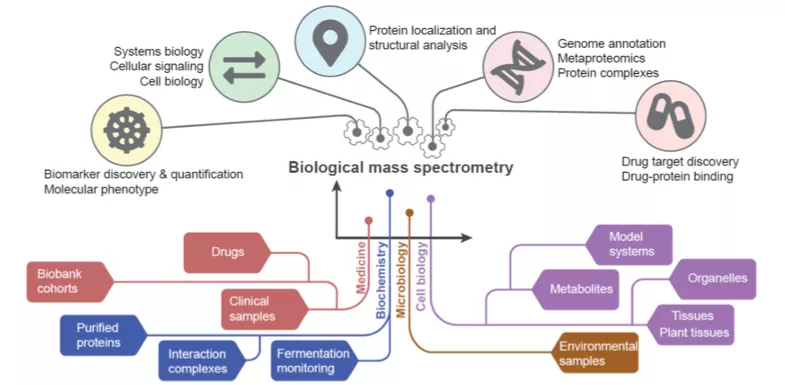
Our facilities provide the opportunity to study protein structure, molecular probes and drug design, system biology and molecular interactions in cells and tissues. Thanks to our world-class research infrastructure, Lund University is well equipped to help lead the way towards future scientific breakthroughs.
Below is a non-exhaustive list of in-house infrastructures that are categorized into four overarching themes:
- bio-imaging
- proteins
- genes and cells
- other resources
Bio-Imaging
Our facilities provide the opportunity to study molecules, cells, organs and entire organisms.
- Correlative Image Processing and Analysis (CIPA)
- LBIC - Lund University Bioimaging Center
- LU-Fold
- MAX IV
- MedMAX a future part of the MAX IV Laboratory in Lund
- Tissue Micro Array Center
- Two-photon microscopy at BMC
Proteins
With the help of various forms of mass spectrometry, synchrotron radiation, protein production & labelling, and bioinformatics, our facilities provide the opportunity to study protein structure and dynamics, molecular probes and drug design. Below you can see some examples of the infrastructure for proteins, available for researchers at Lund University.
- Biacore X100 - Surface Plasmon Resonance
- BioMAX at MAX IV
- BioMS
- CTP - Center for Translational Proteomics
- LP3 - Lund Protein Production Platform
- LU-Fold
- MAX IV
- MESO QuickPlex at MultiPark Platform
Genes and cells
Our facilities provide the opportunity to study molecular probes, system biology and molecular interactions in cells and tissues. Below you can see some examples of the infrastructure for research on genes and cells, available for researchers at Lund University.
- Cell and Gene Therapy Core Facility
- Cellomics and Flow Cytometry Core Facility at MultiPark
- CTG - Center for Translational Genomics
- HistoCore Facility (HCF)
- Life Science Microfluidics
- LUDC Flow Cytometry Core Facility in Malmö
- SpatialOmics@LU
- StemTherapy FACS Core Facility
Other resources
In addition to infrastructures for bioimaging, protein and genes & cells, we also provide other resources e.g., databases, networks and specialized labs.
- CCM - Centre for Comparative Medicine
- Drug Candidate Screening Platform
- EpiHealth Cohort Study
- HALRIC - Hanseatic Life Science Research Infrastructure Consortium
- IQ Biotechnology Platform
- LUBI - Lund University Bioinformatics Infrastructure
- LUPOP (Lund University Population Research Platform)
- MoRe-Lab
- The Scania Metadatabase for Epidemiology
List of infrastructures in alphabetical order
- Biacore X100 - Surface Plasmon Resonance
- BioMAX at MAX IV
- BioMS
- Cell and Gene Therapy Core Facility
- Cellomics and Flow Cytometry Core Facility at MultiPark
- Centre for Comparative Medicine (CCM)
- Correlative Image Processing and Analysis (CIPA)
- CTG - Center for Translational Genomics
- CTP - Center for Translational Proteomics
- Drug Candidate Screening Platform
- EpiHealth Cohort Study
- HistoCore Facility (HCF)
- IQ Biotechnology Platform
- LBIC - Lund University Bioimaging Center
- Life Science Microfluidics
- LP3 - Lund Protein Production Platform
- LU-Fold
- LUBI - Lund University Bioinformatics Infrastructure
- LUDC Flow Cytometry Core Facility in Malmö
- LUPOP (Lund University Population Research Platform)
- MAX IV
- MedMAX a future part of the MAX IV Laboratory in Lund
- MESO QuickPlex at MultiPark Platform
- MoRe-Lab
- The Scania Metadatabase for Epidemiology
- SpatialOmics@LU
- StemTherapy FACS Core Facility
- Tissue Micro Array Center
- Two-photon microscopy at BMC
Biacore X100 - Surface Plasmon Resonance
The Biacore X100 instrument is open for external users to hire after completed course or we can provide a full service.
The Biacore X100 uses the surface plasmon resonance (SPR) technique to monitor molecular interactions in real time. The procedure is as follows; one of the interacting molecules (the ligand) is attached to a surface of a sensor chip, and the other interacting molecule (analyte) is injected and will flow over the sensor chip. If any binding occurs between the ligand and the analyte, the refractive index will change at the sensor surface, that is proportional to the change in mass concentration, and association and dissociation rates of the interaction may be determined.
Read more: Biacore X100- Surface Plasmon Resonance (Lund University Research Portal)
BioMAX at MAX IV
BioMAX is the first X-ray macromolecular crystallography beamline of MAX IV Laboratory. It is a state-of-the-art resource accommodating multiple cutting edge experimental possibilities. The beamline experiment set-up is highly automated, in terms of both sample handling hardware and data analysis.
Due to its extensive energy tunability, BioMAX is an ideal source for de novo phasing using the anomalous signal of heavy elements. The small beam cross-section and parallel beam makes BioMAX suitable for X-ray crystallography using microcrystals and ultra large unit cells.
Read more: BioMAX at MAX IV (MAX IV website)
BioMS
BioMS is a national research infrastructure supported by the Swedish Research Council with a mission to act as a Swedish hub for service, development and coordination of Biological Mass Spectrometry. Currently 40 technical experts, applications specialist and researchers are working in Lund, Stockholm and Gothenburg. The support is organized in two modules of operation, i.e. Quantitative and Structural MS.
Quantitative MS
- Large scale quantitative MS
- Clinical proteomics
- Proteogenomics
- Targeted MS
Structural MS
- Post translational modifications MS
- Interaction MS
- Chemical proteomics
For more information visit our website or contact Lab Manager:
- BioMS website
- sven [dot] kjellstrom [at] med [dot] lu [dot] se (Lab Manager Sven Kjellström (e-mail))
Cell and Gene Therapy Core Facility
The Cell and Gene Therapy core is an open-access, fee-for-service infrastructure supported by Stem Cell Centre and MultiPark. Our services include AAV and LV vector production, cloning, iPS reprogramming, iPS-edits and CRISPR experimental designs. The core is located at BMC B10 and is open to all national academic institutions.
Read more: Cell and Gene Therapy Core Facility (Stem Cell Center website)
Cellomics and Flow Cytometry Core Facility at MultiPark
Screening/Analysis and FACSAria III are open to external users.
The Cellomics Array Scan is a high throughput system for acquisition and screening of Fluorescence Microscopy images, applying automated image analysis of fluorescence intensities in cells, subcellular compartments or small organisms, e.g. zebrafish and nematodes. Our BD FACSAriaIII cell sorter is equipped with three lasers (blue, yellow/green, red) and can analyze up to ten fluorescent channels and sort up to four cell populations simultaneously.
Read more on MulitPark website: FACS platform at MultiPark and Cellomics platform at MultiPark
CCM - Centre for Comparative Medicine
The Centre for Comparative Medicine (CCM) is an infrastructure within the Medical Faculty at Lund University enabling researchers to carry out cutting edge translational research using animal models in Lund and Malmö.
Specialist veterinarians and geneticists are available to advise researchers on aspects of experimental design, ethical permit applications, animal health and husbandry, and advanced genetic engineering techniques. Skilled technical staff provide the daily care for around 35,000 animals (mostly rodents) and maintain the high levels of hygiene necessary to protect the animals. Swedish Animal Welfare regulations and 3R principles (Replacement, Reduction and Refinement) are followed strictly to ensure good animal welfare.
Read more: Centre for Comparative Medicine (CCM)
Correlative Image Processing and Analysis (CIPA)
CIPA is hosted by the Faculty of Medicine and is the Lund University infrastructure for image processing and analysis. Scientific progress heavily relies on comprehending data either by extracting information from images or generating images to understand the data – services provided by CIPA. Addressing challenges in analysis and processing, due to high dimensionality, noisy data, and intricate complexity often extends beyond the expertise of the respective scientists and requires specialized knowledge in image processing and analysis. At Lund University CIPA caters for these much-needed resources.
Read more: CIPA website
CTG - Center for Translational Genomics
CTG is a research infrastructure and technical platform for Next Generation Sequencing (NGS) and other genomics technologies. CTG is part of the SciLifeLab infrastructure. We provide the following services:
- Support in project planning
- Guidelines for DNA and RNA extractions
- Quality control and Library preparation (standard and customized)
- Next Generation Sequencing (DNA and RNA)
- Bioinformatics expertise
Read more: CTG website
CTP - Center for Translational Proteomics
The Center for Translational Proteomics (CTP) was established in 2018 and is financed by the Faculty of Medicine, Lund University and Region Skåne. CTP works closely with the Swedish national infrastructure for biological mass spectrometry (BioMS, Lund Node) supported by the Swedish Research Council. CTP offers expertise and support in small proteomics and protein characterisation projects for scientists at the Faculty of Medicine, Lund University and Region Skåne. CTP helps with study design, mass spectrometry analysis (identification and quantification of proteins) and analysis of the data.
Contact: Charlotte [dot] Welinder [at] med [dot] lu [dot] se (Charlotte[dot]Welinder[at]med[dot]lu[dot]se)
CTP (Lund University Research Portal)
Drug Candidate Screening Platform
The drug candidate screening platform support researchers to identify and develop novel chemical tools to help you interrogate your particular field of interest. We contribute with +30 years of combined experiences from the pharmaceutical industry.
Services provided
- Consultations within medicinal chemistry, chemical biology and drug discovery and development
- Small molecule screening
- assay development/optimization
- screening using in house libraries or from collaborators
- mechanism studies using pathway specific small molecule libraries
- guidance follow-up medicinal and synthetic chemistry
- facilitate project transfer to/collaboration with other infrastructures
More information (MultiPark website)
EpiHealth Cohort Study
The EpiHealth cohort comprises a total of 25,104 individuals from the Swedish cities Uppsala and Malmö in the age groups 45-75 years and was collected during 2010-2016.
Data include lung function (spirometry), inch ECG, blood pressure, body composition, cognitive test (Trial-Making-Test), fasting blood glucose and blood lipids. A questionnaire that includes sociodemographic factors, lifestyle factors, previous illnesses.
A biobank is available for future use and a DNA extraction with GWAS analysis has been carried out throughout the cohort.
Read more: Epihealth Cohort Study (EpiHealth website)
HALRIC
HALRIC aims to be a springboard for innovation capacity in the ÖKS-Hamburg Life Science sector via increased access to and use of cross-border front-end technologies, instruments, expertise, and data handling solutions.
HALRIC is an EU-supported project working to bring more companies, hospitals and academic researchers together in collaborations with:
- The large-scale infrastructures, such as MAX IV & DESY and ESS & European XFEL
- And with cutting-edge complementary infrastructures such as cryoEM, advanced biological MS, and 7Tesla-MR.
HistoCore Facility (HCF)
The HistoCore facility offers services for studying cells and tissue specimens of experimental animal and human origin. The purpose is to facilitate for researchers to connect to the pathology department and obtain well-annotated biobank samples for various omics analyses. The facility strives to interact with and increase the visibility of other similar facilities in the region of Malmö and Lund.
Services provided
The HistoCore facility provides services such as histoprocessing, paraffin embedding, paraffin sectioning, frozen sectioning, construction of tissue microarrays (TMA) and paraffin embedded cell blocks, immunohistochemistry, and extraction of DNA/RNA from human and animal tissues, cells in culture and blood. We also provide services for slide scanning.
Contact: Bjorn [dot] nodin [at] med [dot] lu [dot] se (Bjorn[dot]nodin[at]med[dot]lu[dot]se)
IQ Biotechnology Platform
IQ Biotechnology gives you access to a whole spectrum of services from experimental design, to gene-, protein- and in vivo- experiments and all the way to analysis. IQ Biotechnology do: Imaging and quantification, experimental design, assay development,high throughput experiments, validation, and data analysis:
- Immunochistochemistry
- Light and Electron Microscopy (TEM, SEM and CLEM)
- Cell analysis - BioPlex
- Cytokine - Biacore
- Gene expression - qPCR
Read more: IQ Biotechnology (Lund University Research Portal)
LBIC - Lund University Bioimaging Center
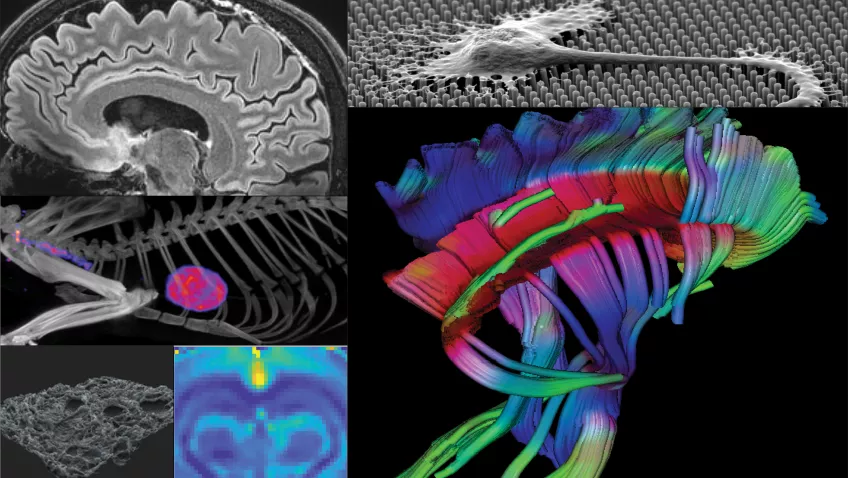
Lund University Bioimaging Centre, LBIC, is a national infrastructure that provides access to a large variety of advanced imaging equipment and cutting-edge bioimaging techniques, ranging from micro to macro. The overall goal is to help lead the way towards future scientific breakthroughs by enabling cutting-edge bioimaging techniques to be part of your research project!
Services
Dedicated and highly experienced staff scientists offer support throughout the entire experimental process – from study design to data handling. LBIC is also offering training, support, workshops and seminars. Our services are available to users from academia, industry and health care and regardless of your particular field of research, you are welcome to seek support from any of our facilities!
Equipment
The infrastructure is built up by several facilities covering all types of bioimaging research:
Microscopy
- Confocal microscopes; wide-field fluorescence microscope and a combined TIRF/STORM (super resolution) microscope
- Light sheet microscope
- Automated live-cell imaging system (also in Biosafety Level 2 laboratory)
- Transmission electron microscope (TEM)
- Scanning electron microscope (SEM)
Preclinical Nuclear Medicine
- μPET/CT
- μSPECT/CT
- Ultra-high resolution μCT
Preclinical MRI
- MRI system, 9.4 Tesla
- Advanced MRI analysis of data
Clinical MRI
- MRI systems, 1,5 and 3 Tesla
The national 7Tesla Facility
- Equipment for brain, knee, wrist, breast and abdominal imaging
- fMRI stimuli and eye tracking
- Clinical devices, such as contrast injector and ventilator
Analysis & Visualization
- Image analysis software packages suitable for both small and large data sets
- Software to transform imaging results into easy-to-use 3D surface models or ray-traced animations
Contact
If you are interested in using our services, please contact the person responsible for the facility of interest.
- Microscopy: Lina Gefors, lina [dot] gefors [at] med [dot] lu [dot] se (lina[dot]gefors[at]med[dot]lu[dot]se) or
Sebastian Wasserström, sebastian [dot] wasserstrom [at] med [dot] lu [dot] se (sebastian[dot]wasserstrom[at]med[dot]lu[dot]se) - Preclinical Nuclear Medicine: Rita Gidlöf, ritha [dot] gidlof [at] med [dot] lu [dot] se (ritha[dot]gidlof[at]med[dot]lu[dot]se)
- Preclinical MRI: Michael Gottschalk, michael [dot] gottschalk [at] med [dot] lu [dot] se (michael[dot]gottschalk[at]med[dot]lu[dot]se)
- Clinical MRI: Pia Sundgren, pia [dot] sundgren [at] med [dot] lu [dot] se (pia[dot]sundgren[at]med[dot]lu[dot]se)
- The National 7 Tesla Facility: Karin Markenroth Bloch, karin [dot] markenroth_bloch [at] med [dot] lu [dot] se (karin[dot]markenroth_bloch[at]med[dot]lu[dot]se)
- Analysis and Visualization: Jonas Ahlstedt, jonas [dot] ahlstedt [at] med [dot] lu [dot] se (jonas[dot]ahlstedt[at]med[dot]lu[dot]se)
For more information about available equipment, services or how to apply please visit the LBIC website.
Life Science Microfluidics
The resource supports development of microfluidic systems. By enabling access to expertise in microfluidic system design and microfabrication facilities, this platform supports researchers with custom development and characterization of microfluidic devices. Our engineers are highly experienced in microfluidics and neuro-microtechnology as well as micromilling and 3D printing of microfluidic system components for many years and available to support microfluidic system development for a multitude of applications.
Equipment and resources
- Clean room facility for basic silicon and glass microfabrication as well as metal film deposition and patterning.
- SU-8 photolithography.
- High resolution direct laser exposure. Glass silicon anodic bonding.
- PDMS microfabrication.
- Precision CNC micromilling.
- Polymer 3D-printing (Fused Filament Fabrication as well as laser induced polymerization).
Services provided
Life Science Microfluidics provides engineering support in research specific development of microfluidic solutions. Support includes device design and manufacturing as well as advice in device assembly and interconnecting to commercial system components (pumps, valves, flow control etc.) for full system functionality.
Contact
Anja Meissner anja [dot] meissner [at] med [dot] lu [dot] se
Thomas Laurell thomas [dot] laurell [at] bme [dot] lth [dot] se
LP3 - Lund Protein Production Platform
A cross-faculty facility for protein production and purification with:
- Cloning - design of a vector construct
- Protein production in bacterial cells and insect cells
- Protein labelling
- Protein purification using chromatography equipment
At LP3 we exchange scientific ideas for dissemination and assimilation of new methodologies for protein production, purification and analysis. We also offer research training and development of skills in experimental protein science for PhD students and postdocs.
Read more: LP3 - Lund Protein Production Platform (Lund University Research Portal)
LU-Fold - a protein structure prediction facility
LU-Fold is a Lund University-based facility for helping researchers predict protein structures of interest using the cutting-edge method AlphaFold2 (Nature Methods method of the year, 2021). LU-Fold specialises in high-throughput prediction of protein complexes to predict novel protein-protein interactions.
More information about LU-Fold
LUBI - Lund University Bioinformatics Infrastructure
Lund University Bioinformatics Infrastructure (LUBI) was established in the end of 2020 as an open research infrastructure/network for bioinformatics, accessible to all researchers and students interested in bioinformatics. The goal for LUBI is to strengthen and coordinate bioinformatics at Lund University. LUBI aims to increase the visibility of bioinformatics resources, organize seminars and workshops, and promote knowledge exchange.
Read more: LUBI website
LUDC Flow Cytometry Core Facility in Malmö
The Flow Cytometry Core Facility at the Lund University Diabetes Centre (LUDC) provides instrumentation and technical assistance to perform flow cytometric analysis and operator based cell sorting service to all Lund University scientists as well as scientists from other institutes and industry.
Our facility houses on cell sorter (BD Aria Fusion), two analysers (CytoFlex and Gallios, both from Beckman Coulter) and a multiplex analyser (Bio-Rad).
Flow Cytometry Core Facility (LUDC website)
LUPOP (Lund University Population Research Platform)
LUPOP is an open resource available for all researchers at Lund University. LUPOP provides support for population based research, such as advice on study design and scientific approaches, applications for funding and data access within population research, and ethical and legal issues. Through the website www.lupop.lu.se, researchers and students can keep updated with the latest within population based research. The website can also be used by researchers at Lund University to provide information to colleagues, students, media, and the public.
Read more: LUPOP website
MAX IV
The MAX IV Laboratory serves 1000 users per year and deliver science in a broad range of fields including structure and dynamics of proteins in solution, protein crystallography and biomedical imaging. Since summer 2016 the new world leadning synchotron facility, MAX IV is up and running.
Read more: MAX IV website
MedMAX a future part of the MAX IV Laboratory in Lund
MedMAX – a future beamline inside MAX IV, which will help us to study events and processes in tissues and materials using accelerators producing x-rays of very high intensity. MedMAX will be a super-eye capable of helping us understand biomedical problems. With its help, we can find new ways of discovering, diagnosing and treating diseases such as cancer, diabetes and osteoporosis.
Read more about MAX IV: MAX IV website
MESO QuickPlex at MultiPark Platform
The MESO QuickPlex is a multiplexing plate reader using high-performance, electrochemiluminescence (ECL) immunoassays for detecting bio markers.
It is quick (90 seconds for one plate) and can handle many samples at the same time. The ECL system allows for detection with high sensitivity and wide dynamic range, which in combination with multi-array technology enables accurate measurements of multiple analytes in low sample volume. The instrument is easy to learn and use. There is more than 600 immunoassays for a range of applications including immunology, inflammation, oncology, neurobiology, and toxicology, in various assay formats.
Read more: MESO QuickPlex at MultiPark Platform (MultiPark website)
MoRe-Lab
The Movement and Reality Lab (MoRe-Lab) is an infrastructure designed as a hub for collaborations between University Faculties, municipalities, health care networks, sports clubs and private industry.
MoRe-Lab houses a live-in instrumented home environment platform (Reality platform) located next to state-of-the-art human movement analysis platforms (Movement platform), thus enabling the study of complex interaction between physical function, behaviour, cognition, and activity in different standardised or real environments.
MoRe-Lab is a unique infrastructure in Sweden and arguably in the World. This facility is not discipline-specific, but rather strives to support interdisciplinary collaboration among traditional health sciences disciplines and any other discipline studying aspects relevant for health.
Read more: MoRe-Lab website
The Scania Metadatabase for Epidemiology
The Scania Metadatabase of Epidemiology is a catalogue with a description of several of the epidemiological studies that have been collected within Lund University and Region Skåne. The catalogue currently comprises 50 studies including a total of more than 670 000 study participants, as well as biological samples from about 20 studies.
The expectation is that the metadatabase will be used as an inspiration for new research and open up new research partnerships by visualizing the research groups and their publications.
Contact: jonas [dot] bjork [at] med [dot] lu [dot] se
SpatialOmics@LU
The SpatialOmics@LU facility at the department of Immunotechnology provides hi-plex, spatial expression analysis with the GeoMx DSP instrument. Investigate 1800+ RNA or up to 80 protein targets in specific regions of tissue sections defined by the morphology. The facility also offers targeted gene expression analysis of up to 800 targets with the nCounter system.
Read more: SpatialOmics@LU website
The Department of Immunotechnology has several other infrastructures. Read more: Infrastructures at the Department of Immunotechnology (LTH website)
StemTherapy FACS Core Facility
Flow cytometry is one of the major methods used to analyze and sub-fractionate cells. It has major advantages such as the simultaneous detection of a wide range of parameters that enables the understanding of diversity among seemingly similar cells. The ability to enumerate large numbers of viable cells in a relatively short period of time provides experiment with reliability. At the Stem Therapy FACS Core Facility we have three cell FACS Aria sorters, that allow detection of up to 16 fluorescent probes, as well as two analysis machines: Fortessa and LSRII. We can help you through the whole process, from design of the experiment to the running of the experiment.
- Assisted Cell sorting
- Assisted FACS Analysis
- Advice and supervision of users during experiments
Advice for experimental design and developing new protocols
Read more: FACS on StemTherapy website
Tissue Micro Array Center
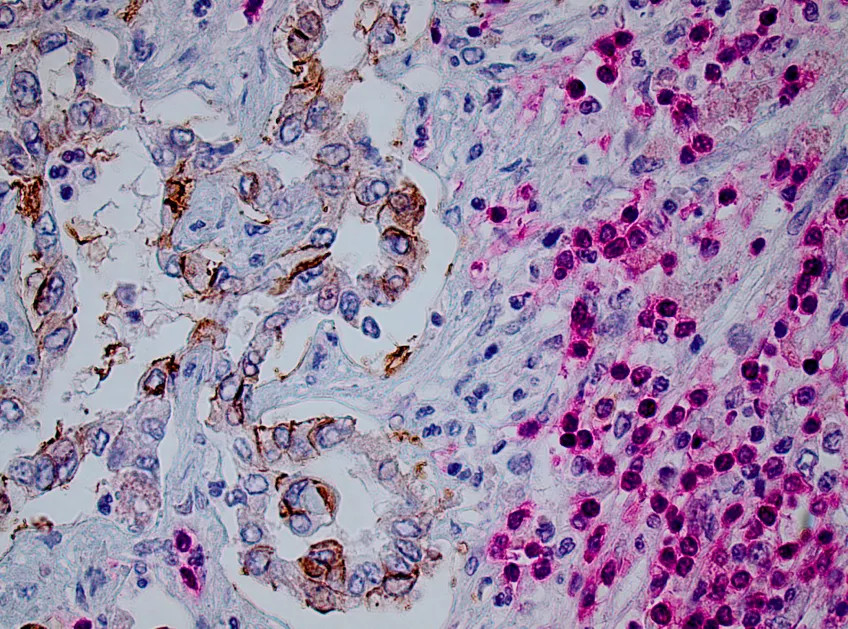
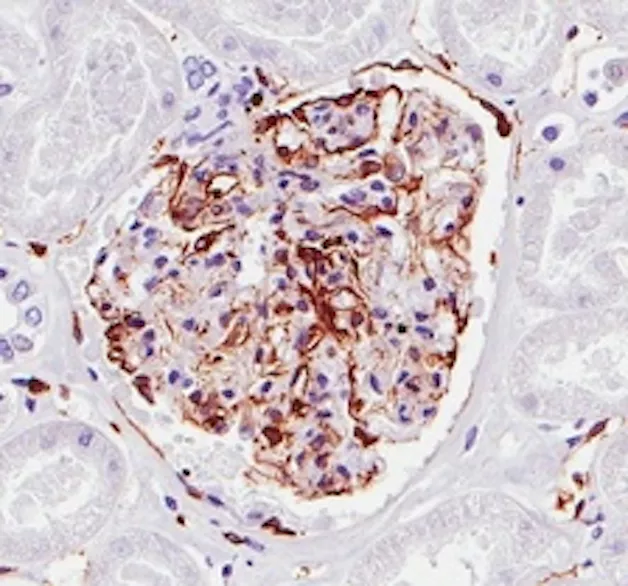
The TMA-center offers high quality immunohistochemistry staining of tissue. Sample preparation and processing, embedding, sectioning, staining and scanning of the stained slides. We provides both single staining (one protein) and double staining (two proteins) performed with an Autostainer (Agilent/DAKO) automated system. The stained slides are scanned using an Aperio scanner.
The TMA-center also assembles TMA-blocks. A TMA block (Tissue Micro Array) is a block consisting of small cores from up to 96 different samples/cancers/tissues all organized in one block.
Contact: kristina [dot] ekstrom-holka [at] skane [dot] se (Kristina Ekström-Holka)
Two-photon microscopy at BMC
This platform allows imaging of tissues with very high temporal and spatial resolution. It consists of two microscopy stations: an ‘in vivo’ setup is used to image dynamic processes in the life mouse and an ‘ex vivo’ system designed to image thick live preparations, such as tissues that are not accessible to ‘in vivo’ microscopy. Both systems are available for LU and external users.
Read more: Two-photon-microscopy (MultiPark website)
Contact
Kajsa M Paulsson
Coordinator Research Infrastructure
e-mail: kajsa_m [dot] paulsson [at] med [dot] lu [dot] se (kajsa_m[dot]paulsson[at]med[dot]lu[dot]se)